MOPS
MOPS - Mathematical Optimization of Parkinson’s Scale
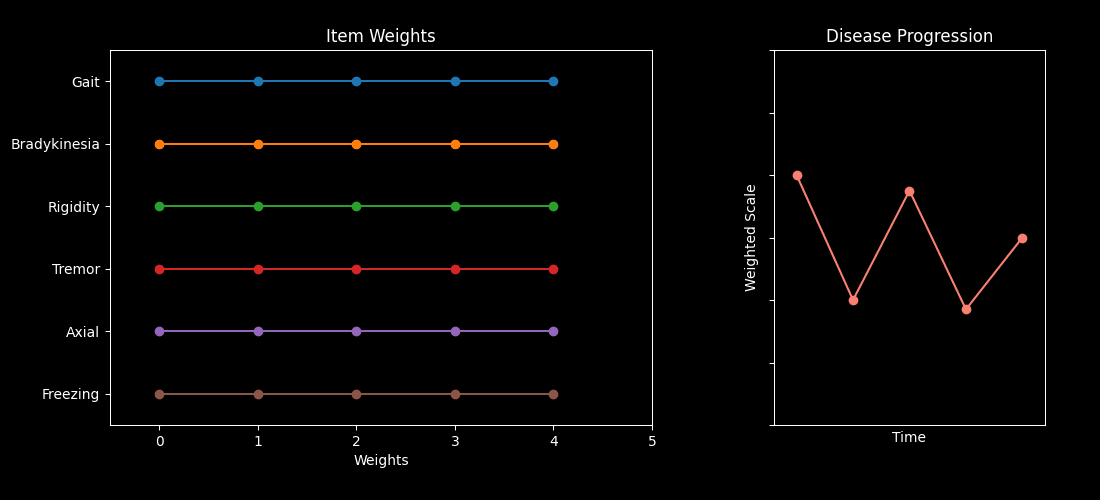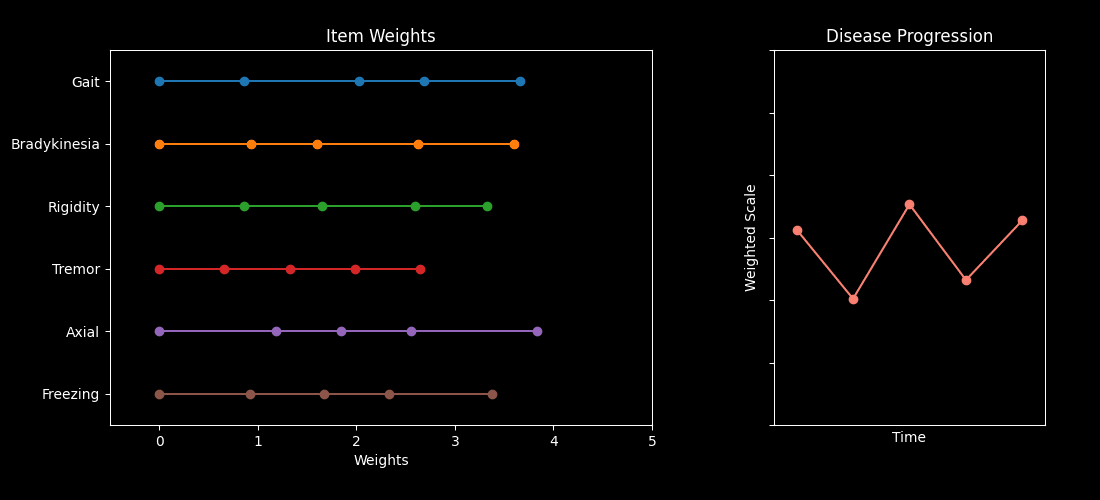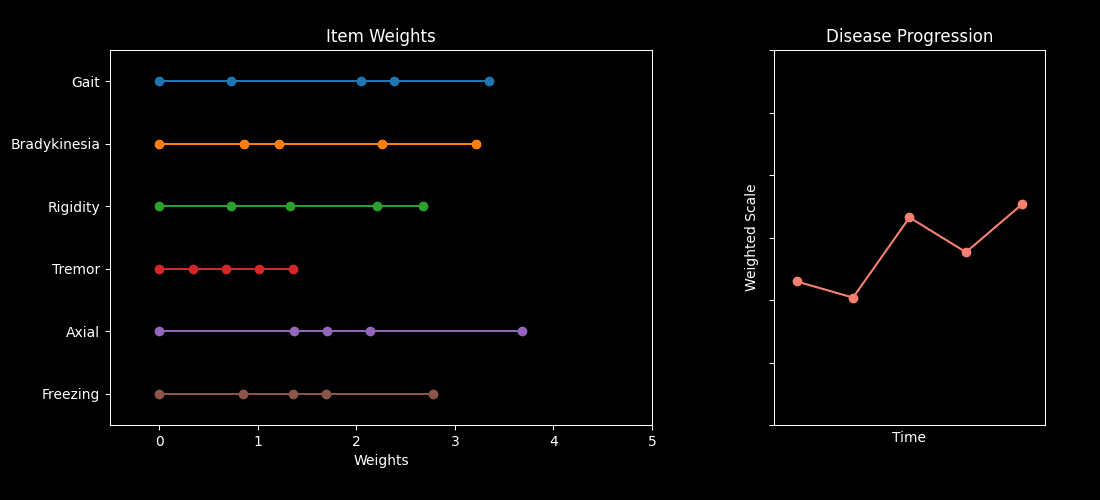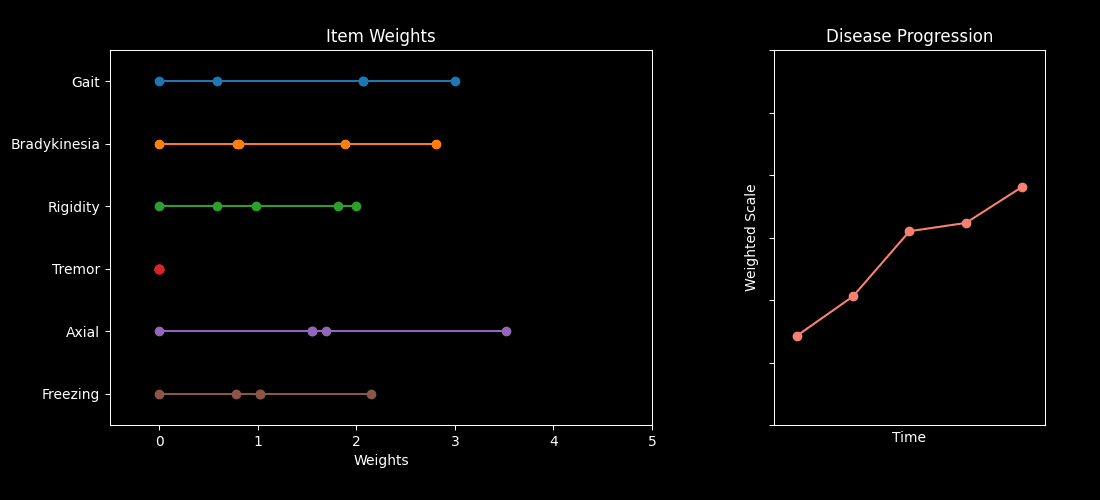
This repository contains code accompanying the paper “Optimizing Parkinson’s Disease progression scales using computational methods.” https://www.nature.com/articles/s41531-026-01259-1
The main goal is to demonstrate how to learn data-driven weights for items in standard Parkinson’s Disease (PD) assessments — such as the MDS-UPDRS and MoCA — so the resulting composite scores better capture disease progression.
Table of Contents
- Repository Structure
- Installation & Dependencies
- Data Requirements
- Usage
- Online Tool
- License
- Contact
Repository Structure
.
├── data/
│ └── PPMI/ # Where to place PPMI data files
├── optimizers/
│ ├── <various_optimizer_files>.py # Implementations of the optimization methods
│ └── weights/ # Directory where generated CSV files of item weights are saved
├── pipeline.ipynb # Main flow: data prep, optimization, evaluation, visualization
├── data_preparation.py # Data filtering and encoding logic
├── requirements.txt # Dependencies for Python environment
├── self_report_short.html # A demo of a short self-reported questionnaire
└── LICENSE # GNU General Public License
Installation & Dependencies
- Clone this repository to your machine:
git clone https://github.com/Shamir-Lab/MOPS.git cd MOPS - Create and activate a Python virtual environment (recommended):
python -m venv venv source venv/bin/activate # On Windows: venv\Scripts\activate - Install requirements:
pip install -r requirements.txt
Gurobi Installation
Several optimizers in optimizers/ rely on the Gurobi solver for integer or mixed integer programming.
To use these formulations:
- Obtain a Gurobi license: Gurobi offers free academic licenses for qualified institutions.
- Install Gurobi: Follow instructions on Gurobi’s website to download and install the Gurobi engine and Python bindings.
- Configure: Ensure your
GUROBI_HOMEandPATH/LD_LIBRARY_PATHare set appropriately (platform-specific instructions are available on their site).
Data Requirements
This code uses the Parkinson’s Progression Markers Initiative (PPMI) data, which is not distributed here. To replicate our analysis:
- Register at the PPMI website and request access to download the clinical datasets.
- Download the following directories: motor and non motor assessment, subject characteristics and medical history.
- Place them under
data/PPMIin this repository.
Usage
- Open JupyterLab or Jupyter Notebook, activate your virtual environment, and navigate to the repository folder.
- Open
pipeline.ipynb - Run the cells in order
Online Tool
Based on our results, we’ve created an online self-reported questionnaire that achieves good consistency with only 11 simple questions.
Tool is available here.
License
This project is licensed under the terms of the GNU General Public License v3.0. You are free to use, modify, and distribute this code under the conditions detailed in the license, which requires that derivative works also be distributed under the same license.
Contact
If you have any questions or suggestions regarding this repository or the associated paper:
- Email: [assafb2@mail.tau.ac.il]
- GitHub issues: Feel free to open an issue in this repository for bugs or feature requests.